JOBIM 2012
Journées Ouvertes en Biologie, Informatique et Mathématiques
3 - 6 juillet 2012
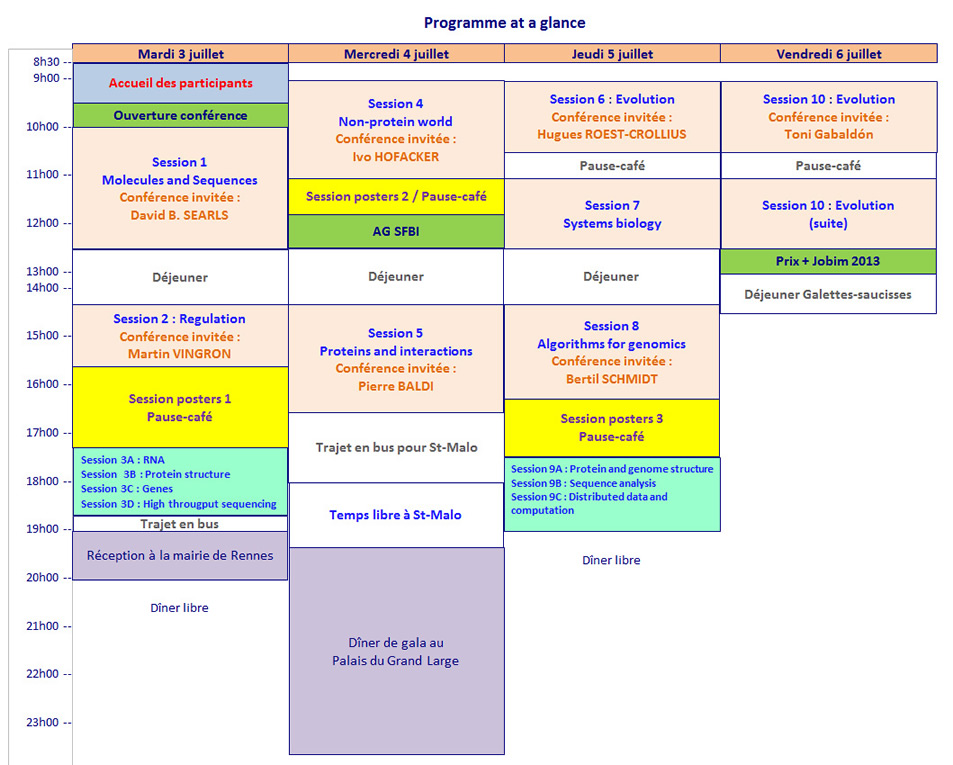
 programme détaillé JOBIM 2012
programme détaillé JOBIM 2012
Mardi 3 juillet |
| 8h30 - 9h45 | Accueil des participants |
| 9h45 - 10h00 | Ouverture conférence |
| 10h00 - 12h30 | Session1 - Molecules and Sequences |
Conférence invitée : David B. Searls - Molecules, Languages, and Automata. |
|
| Expressive Pattern Matching with Logol. Application to the Modelling of -1 Ribosomal Frameshift events. Catherine Belleannée, Olivier Sallou and Jacques Nicolas |
|
| SA-Mot: a new web server for the extraction of motifs of interest from protein loops. Leslie Regad, Juliette Martin, Colette Geneix and Anne-Claude Camproux |
|
| The sequence space of G-protein-coupled receptors: Implications for molecular modeling. Jean-Michel Becu, Julien Pele, Hervé Abdi and Marie Chabbert |
| 12h30 - 14h15 | Déjeuner |
| 14h15 - 15h30 | Session 2 - Regulation |
Conférence invitée : Martin Vingron - Computational Regulatory Genomics. |
|
| Institut Français de Bioinformatique Jean-François Gibrat |
| 15h30 - 17h15 | Session posters 1 / pause-café |
| 17h15 - 18h30 | Sessions parallèles |
Session 3-A : RNA |
|
| Development of a pipeline for Scyliorhinus canicula miRNA identification from NGS data. Wilfrid Carré, Pierre Pericard, Christophe Caron, Erwan Corre and Sylvie Mazan |
|
| BoostSVM: A miRNA classifier with high accuracy using boosting SVM. Van Du Tran, Benjamin Zerath, Sebastien Tempel, Farida Zehraoui and Fariza Tahi |
|
| miRNAFold: A fast ab-initio method for searching for miRNA precursors in whole genomes. Sébastien Tempel and Fariza Tahi |
Session 3-B : Protein structure |
|
| A Modular Network Analysis (MONETA) of Protein Structures for Exploring and Predicting Allosteric Communication. Elodie Laine, Christian Auclair and Luba Tchertanov |
|
| Impact of the D802V Mutation on the Structure and Dynamics of the CSF-1R Tyrosine Kinase Receptor. Priscila Da Silva Figueiredo Celestino, Elodie Laine, Pedro Geraldo Pascutti and Luba Tchertanov |
|
| Structural determinants of Raltegravir specific recognition by the HIV-1 Integrase. Rohit Arora and Luba Tchertanov |
Session 3-C : Genes |
|
| Importance of family size and function in the fate of duplicated genes in the protein-protein interactome of Arabidopsis thaliana. Justin Whalley, Etienne Birmelé and Carène Rizzon |
|
| GO2PUB PubMed Query Tool Based on Semantic Expansion of Gene Ontology Terms, a Lipid Metabolism Case Study. Charles Bettembourg, Christian Diot, Anita Burgun and Olivier Dameron |
|
| Retrospective Analysis of a Gene by Mining Texts: The Hepcidin Gene Use-Case. Fouzia Moussouni, Bertrand Ameline De Cadeville, Ulf Leser and Olivier Loréal |
Session 3-D : High througput sequencing |
|
| Rapid Prioritization and Annotation of Variants from Human Re-sequencing Studies. Elodie Dubus, D Richards, R Flannery, A Kramer, J Lerman et A Kutchma (Ingenuity Systems) |
|
| Extracting relevant information from UHTS data. Patricia Otten-Hernandez, L Baerlocher , J Prados , N González , W Baroni , M Osteras et L Farinelli (Fasteris) |
|
| KLAST: high-performance sequence similarity search tool. Erwan Drezen, Alain Meil, Dominique Lavenier et Patrick Durand (Korilog & Inria/IRISA GenScale team) |
| 19h00 - 20h00 | Réception à la maire de Rennes |
| 20h00 | Dîner libre |
Mercredi 4 juillet |
| 9h00 - 11h00 | Session 4 - Non-protein world |
Conférence invitée : Ivo Hofacker - RNA Structures, Interactions, and Folding Kinetics |
|
| Computational detection and expression profiling of conserved long non-coding RNAs in the domestic dog. Thomas Derrien, Amaury Vaysse, Benoit Hennuy, Wouter Coppieters, Benoit Hedan, Catherine Andre and Christophe Hitte |
|
| NRPS toolbox for the discovery of new nonribosomal peptides and synthetases. Maude Pupin, Malika Smaïl-Tabbone, Philippe Jacques, Marie-Dominique Devignes and Valerie Leclère |
| 11h00 - 11h45 | Session Posters 2 / Pause-café |
| 11h45 - 12h30 | AG SFBI |
| 12h30 - 14h15 | Déjeuner |
| 14h15 - 16h30 | Session 5 - Proteins and interactions |
Conférence invitée : Pierre Baldi -Machine Learning Approaches in Proteomics |
|
| Protein-protein interaction network inference with semi-supervised Output Kernel Regression. Celine Brouard, Marie Szafranski and Florence D'Alché-Buc |
|
| Improving gene signatures by the identification of differentially expressed modules in molecular networks : a local-score approach. Marine Jeanmougin, Christophe Ambroise, Matthieu Bouaziz and Mickaël Guedj |
|
| Présentation www.bioinfo-fr.net. Yoann Mouscaz |
| 16h30 | Départ pour St-Malo |
| 18h00 - 19h15 | Temps libre à St-Malo |
| 19h15 - 00h00 | Dîner de gala au Palais du Grand Large |
Jeudi 5 juillet |
| 9h00 - 10h30 | Session 6 - Evolution |
Conférence invitée : Hugues Roest Crollius - The 4th dimension of Biology. A historical perspective of biological processes. |
|
| Non-adaptive expansion of gene families. Param-Priya Singh, Severine Affeldt and Herve Isambert |
| 10h30 - 11h00 | Pause café |
| 11h00 - 12h30 | Session 7 - Systems biology |
| Logical modelling of cellular decision processes with GINsim. Claudine Chaouiya, Aurélien Naldi, Lionel Spinelli, Pedro T. Monteiro, Duncan Berenguier, Luca Grieco, Abibatou Mbodj, Samuel Collombet, Anna Niarakis, Laurent Tichit, Elisabeth Remy and Denis Thieffry |
|
| Using large scale knowledge databases about reactions and regulations to find key regulators of sets of genes. Pierre Blavy, Florence Gondret, Sandrine Lagarrigue, Jaap van Milgen and Anne Siegel |
|
| Using Mutual Information and Answer Set Programming to refine PWM based transcription regulation network. Andres Aravena, Carito Guziolowski, Anne Siegel and Alejandro Maass |
| 12h30 - 14h15 | Déjeuner |
| 14h15 - 16h15 | Session 8 - Algorithms for genomics |
Conférence invitée : Bertil Schmidt - Scalable Algorithms and Tools for Biological Sequence Analysis. |
|
| Mapping Reads on a Genomic Sequence: a Practical Comparative Analysis. Sophie Schbath, Véronique Martin, Matthias Zytnicki, Julien Fayolle, Valentin Loux and Jean-François Gibrat |
|
| SortMeRNA: a new software to filter total RNA for metatranscriptomic or RNA analysis. Evguenia Kopylova, Laurent Noé and Hélène Touzet |
|
| Présentation Jebif |
| 16h15 - 17h30 | Session posters 3 / pause-café |
| 17h30 - 19h00 | Sessions parallèles |
Session 9-A : Protein and genome structure |
|
| CSA: Comparaison compréhensive d'alignement de paires de structures de protéines. Inken Wohlers, Noël Malod-Dognin, Rumen Andonov and Gunnar Klau |
|
| A Novel Approach of Spatial Motif Extraction to Classify Protein Structures. Rabie Saidi, Wajdi Dhifli, Mondher Maddouri and Engelbert Mephu Nguifo |
|
| On the Complexity of two Problems on Orientations of Mixed Graphs. Hafedh Mohamed Babou, Guillaume Fertin and Irena Rusu |
Session 9-B : Sequence analysis |
|
| Fast Homology Search Using Domain-Architecture Alignment. Nicolas Terrapon, Sonja Grath, January Weiner, Andrew D. Moore and Erich Bornberg-Bauer |
|
| Fast estimation of posterior probabilities in change-point models through a constrained hidden Markov model. The Minh Luong, Yves Rozenholc and Gregory Nuel |
|
| Detecting Outliers in HMM modeling through Relative Entropy with Applications to Change-Point Detection. Vittorio Perduca and Gregory Nuel |
Session 9-C : Distributed data and computation |
|
| Towards a Distributed Infrastructure for Bioinformatics: French GRISBI Perspective. Christophe Blanchet, Clément Gauthey, Christophe Caron, Olivier Collin, Stéphane Delmotte, Tiphaine Martin, Simon Penel, Aurélien Roult, Franck Samson and Bruno Spataro |
|
| Parallel Storage: Addressing the Big Data Challenge. Sven Neirynck (Panasas) |
|
| Plus de simulations pour des résultats encore plus rapide avec les GPU NVIDIA. Jean-Christophe Baratault (NVIDIA - Tesla Sales EMEA) |
| 20h00 | Dîner libre |
Vendredi 6 juillet |
| 9h00 - 10h30 | Session 10 - Evolution |
Conférence invitée : Toni Gabaldón - Phylogenomics in the light of ever-growing sequencing data. |
|
| Evolution of conjugation and type IV secretion systems. Julien Guglielmini, Fernando de La Cruz and Eduardo P.C. Rocha |
|
| Présentation DE^CODAGE Philippe Leroy |
| 10h30 - 11h00 | Pause-café |
| 11h00 - 12h30 | Session 10 - Evolution (suite) |
| EvoluCode: an original view of Human Systems Evolution. Benjamin Linard, Olivier Poch and Julie Thompson |
|
| Haplotype-based method for detecting regions under selection in the domestic dog. Hillel Jean-Baptiste-Adolphe, Mathieu Emily, Amaury Vaysse, Catherine André and Christophe Hitte |
|
| GECA: Gene Evolution/Conservation Analysis tool for Eukaryotic gene families. Nizar Fawal, Bruno Savelli, Christophe Dunand and Catherine Mathé |
| 12h30 - 13h00 | Prix + Jobim 2013 |
| 13h00 - 14h30 | Déjeuner Galettes-saucisses |